Using filesets for data processing
What is the idea?
Imagine you have a big dataset consisting of many files containing observations (e.g. images or satellite data). The files cover certain time periods and are bundled into subdirectories. See Fig.19 for an example.
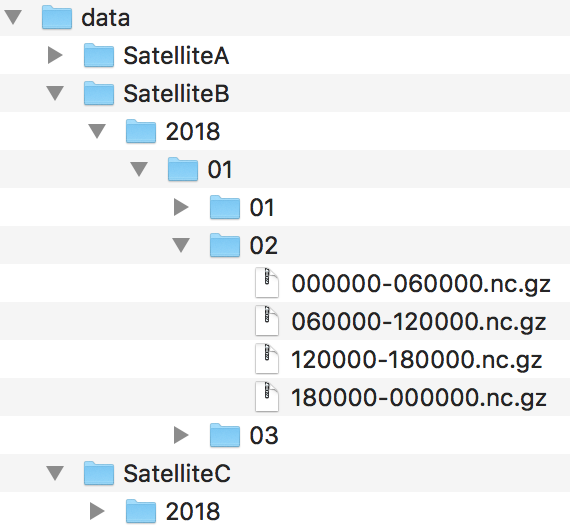
Fig. 19 Example of filesets
All files of Satellite B are located in subdirectories which contain temporal information in their names (year, month, day, etc.).
Typical tasks to analyze this dataset would include iterating over those
files, finding those that cover a certain time period, reading them, applying
functions on their content and eventually adding files with new data to this
dataset. So, how to find all files in a time period? You could start by writing
nested for loops and using python’s glob function. Normally, such solutions
requires time to implement, are error-prone and are not portable to other
filesets with different structures. Hence, save your time/energy/nerves and
simply use the FileSet class.
Hint
NOTE: The code in the jupyter notebooks is old and needs to be updated!
If you want to run the code from this tutorial on your machine as well,
download
spareice_tutorials.zip and
unzip it. You can find the code examples for this tutorial in the jupyter
notebook file dataset.ipynb. You will need the jupyter engine for this.
Find Files
We stick to our example from above and want to find all files from our Satellite B dataset between two dates. To do this, we have to create a FileSet object with the path to our files:
1# Import the FileSet class from the typhon module.
2from typhon.files import FileSet
3
4# Define a fileset object pointing to the files
5# of the Satellite B
6b_fileset = FileSet(
7 path="data/SatelliteB/{year}-{month}-{day}/"
8 "{hour}{minute}{second}-{end_hour}{end_minute}{end_second}.nc"
9)
Nothing interesting happens so far. We imported the FileSet class from the typhon module, created a FileSet object and told it where to find its files. These words surrounded by braces (e.g. “{year}”) are called placeholders. They work like regular expressions and generalize the path, so we need not give explicit paths that point to each file directly. The FileSet object can fill those placeholders by itself when searching for files. Let’s see it in action:
# Find all files (but only print the first one)
for file in b_fileset:
print(repr(file))
1.../data/SatelliteB/2018-01-01/000000-050000.nc
2.../data/SatelliteB/2018-01-01/050000-100000.nc
3.../data/SatelliteB/2018-01-01/100000-150000.nc
4.../data/SatelliteB/2018-01-01/150000-200000.nc
5.../data/SatelliteB/2018-01-01/200000-010000.nc
6.../data/SatelliteB/2018-01-02/010000-060000.nc
7.../data/SatelliteB/2018-01-02/060000-110000.nc
8.../data/SatelliteB/2018-01-02/110000-160000.nc
9.../data/SatelliteB/2018-01-02/160000-210000.nc
10.../data/SatelliteB/2018-01-02/210000-020000.nc
11.../data/SatelliteB/2018-01-03/060000-120000.nc
If we want to have only files from a certain time period, we can use the
find() method with start and end parameter:
# Find all files in a certain time period
for file in b_fileset.find("2018-01-01", "2018-01-01 12:00:00"):
print(file)
.../data/SatelliteB/2018/01/01/000000-050000.nc
.../data/SatelliteB/2018/01/01/050000-100000.nc
In both examples from above, we yield a
FileInfo object in the file
variable. The FileInfo object has three attributes: path, times and attr.
Let’s have a look at them:
print("Type:", type(file))
print("Path:", file.path)
print("Times:", file.times)
print("Attributes", file.attr)
Type: <class 'typhon.files.handlers.common.FileInfo'>
Path: .../data/SatelliteB/2018-01-03/060000-120000.nc
Times: [datetime.datetime(2018, 1, 3, 6, 0), datetime.datetime(2018, 1, 3, 12, 0)]
Attributes: {}
The path returns the path to the file and times is a list with two datetime objects: the start and end time of the file. They are retrieved by the placeholders that were used in the path argument of the Dataset object. But what is about attr and why is it an empty dictionary? Additionally to the temporal placeholders (such as {year}, etc.), which are converted into start and end datetime objects, you can define own placeholders. For example, let’s make a placeholder out of the satellite name:
1# The same dataset as before but with one additional placeholder in the
2# path:
3fileset = FileSet(
4 path="data/{satname}/{year}-{month}-{day}/"
5 "{hour}{minute}{second}-{end_hour}{end_minute}{end_second}.nc"
6)
7
8for file in fileset.find("2018-01-01", "2018-01-02"):
9 print("Path:", file.path)
10 print("Attributes", file.attr)
1Path: .../data/SatelliteA/2018-01-01/000000-040000.nc
2Attributes {'satname': 'SatelliteA'}
3Path: .../data/SatelliteB/2018-01-01/000000-050000.nc
4Attributes {'satname': 'SatelliteB'}
5Path: .../data/SatelliteA/2018-01-01/040000-080000.nc
6Attributes {'satname': 'SatelliteA'}
7Path: .../data/SatelliteB/2018-01-01/050000-100000.nc
8Attributes {'satname': 'SatelliteB'}
9Path: .../data/SatelliteA/2018-01-01/080000-120000.nc
10Attributes {'satname': 'SatelliteA'}
11Path: .../data/SatelliteB/2018-01-01/100000-150000.nc
12Attributes {'satname': 'SatelliteB'}
13Path: .../data/SatelliteA/2018-01-01/120000-160000.nc
14Attributes {'satname': 'SatelliteA'}
15Path: .../data/SatelliteB/2018-01-01/150000-200000.nc
16Attributes {'satname': 'SatelliteB'}
17Path: .../data/SatelliteA/2018-01-01/160000-200000.nc
18Attributes {'satname': 'SatelliteA'}
19Path: .../data/SatelliteA/2018-01-01/200000-000000.nc
20Attributes {'satname': 'SatelliteA'}
21Path: .../data/SatelliteB/2018-01-01/200000-010000.nc
22Attributes {'satname': 'SatelliteB'}
As we can see, we are able to find the data from Satellite A as well because
it has the same subdirectory structure as Satellite B. The placeholder
satname - per default interpreted as wildcard - was filled by Dataset
automatically and returned in attr. This could be useful if we want to
process our files and we need to know from which satellite they came from. We
can apply a filter on this placeholder when using
find():
filters = {"satname": "SatelliteA"}
for file in fileset.find("2018-01-01", "2018-01-02", filters=filters):
print("Path:", file.path)
print(" Attributes", file.attr)
This finds only the files which placeholder satname is SatelliteA. We can also set it to a regular expression. If we want to apply our filter as a black list, i.e. we want to skip all files which placeholders contain certain values, we can add a ! before the placeholder name.
# This finds all files which satname is not SatelliteA
filters = {"!satname": "SatelliteA"}
We can also set a placeholder permanently to our favourite regular expression
(e.g. if you want to call find()
multiple times). Use
set_placeholders() for this:
fileset.set_placeholders(satname="\w+?B")
Which results that we only find satellites which name ends with B. If you want to find out more about placeholders, have a look at this section.
Read and Create Files
Handling common file formats
Well, it is nice to find all files from one fileset. But we also want to open them and read their content. For doing this, we could use our found FileInfo objects as file argument for python’s open builtin function:
for file in b_fileset.find("2018-01-01", "2018-01-02"):
with open(file, "rb") as f:
# This returns a lot of byte strings:
print(f.readline())
Okay, this may be not very practical for netCDF files since it just returns a lot of byte strings. Of course, we could use the python-netcdf module for reading the files but our FileSet object provides a much easier way:
data = b_fileset["2018-01-01"]
print(data)
1<xarray.Dataset>
2Dimensions: (time: 15)
3Coordinates:
4 * time (time) datetime64[ns] 2018-01-01 2018-01-01T00:20:00 ...
5Data variables:
6 lat (time) float64 ...
7 lon (time) float64 ...
8 data (time) float64 ...
This found a file that is the closest to 2018-01-01 and decompressed it (if it was compressed by using zip, gzip or other common compression standards). Afterwards it loaded its content into an xarray.Dataset object (kind of sophisticated dictionary that holds numpy arrays; have a look at xarray). And all this by using only one single expression! We can also read all files from a time period:
1# Find files from 2018-01-01 to 2018-01-01 and load them into
2# numpy arrays
3data = b_fileset["2018-01-01":"2018-01-02"]
4
5# data is now a list of xr.Dataset objects.
What if we want to create a new file for our FileSet? How does this work? It is as simple as reading them. Create your data object first and then pass it to the FileSet:
1import xarray as xr
2
3# Create a xr.Dataset which holds data in form of numpy arrays:
4data = xr.Dataset()
5data['lat'] = 'time', 90 * np.sin(np.linspace(0, 6.28, 7))
6data['lon'] = 'time', np.linspace(-180, 180, 7)
7data['data'] = data['lat'] * 2 + np.random.randn(7)
8data["time"] = pd.date_range(
9 "2018-01-03 06:00:00", "2018-01-03 12:00:00", freq="h"
10)
11
12# Save this xr.Dataset object to a file that belongs to our fileset. The
13# given timestamps indicate the time coverage of the file:
14b_fileset["2018-01-03 06:00:00":"2018-01-03 12:00:00"] = data
If we look now in our dataset directory, we find a new file called data/SatelliteB/2018-01-03/060000-120000.nc. We can see its content with a netCDF viewer (e.g. panoply). So our FileSet object took our xarray.Dataset and put it into a netCDF file automatically. The FileSet object tries to detect from the path suffix the format of the files. This works for netCDF files (*.nc) and for CSV files (*.txt, *.asc or *.csv).
Handling other file formats
What happens with files in CSV format but with a different filename suffix? Or
with other file formats, e.g. such as from CloudSat instruments? Can the
FileSet read and write them as well? Yes, it can. But it is going to need some
help of us before doing so. To understand this better, we have to be honest:
the FileSet object cannot do very much; it simply finds files and compress /
decompress them if necessary. However, to read or create files, it trusts a
file handler and let it do the format-specific work. A file handler is an
object, which knows everything about a certain file format and hence can read
it or use it to write a new file. The FileSet object automatically tries to
find an adequate file handler according to the filename suffix. Hence, it knew
that our files from Satellite B (with the suffix .nc) have to be opened
with the
NetCDF4 file handler.
If we want to use another file handler, we can set the file handler by ourselves. Let’s demonstrate this by using another fileset, e.g. data from Satellite C. Its structure looks like this:
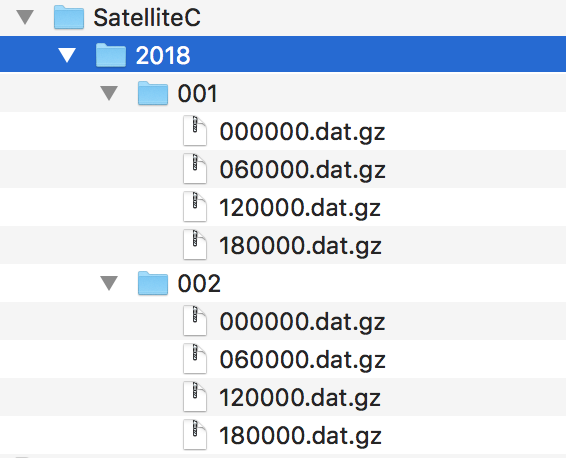
Fig. 20 Files of Satellite C
The files are stored in a different directory structure and are in CSV format. Instead of having subdirectories with month and day, we now have subdirectories with the so-called day-of-year (all days since the start of the year). Do not worry, the FileSet object can handle this structure without any problems with the temporal placeholder doy:
1c_fileset = FileSet(
2 path="data/SatelliteC/{year}-{doy}/{hour}{minute}{second}.dat",
3)
4
5for file in c_dataset.find("2018-01-01", "2018-01-02"):
6 print(file)
.../data/SatelliteC/2018-001/000000.dat
Start: 2018-01-01 00:00:00
End: 2018-01-01 00:00:00
...
But if we try to open one of the files, the following happens:
data = c_dataset["2018-01-01"]
1---------------------------------------------------------------------------
2NoHandlerError Traceback (most recent call last)
3...
4
5NoHandlerError: Could not read '.../data/SatelliteC/2018-001/000000.dat'!
6 I do not know which file handler to use. Set one by yourself.
It cannot open the file because it could not retrieve a file handler from the
filename suffix. Let’s help the Dataset object by setting its file handler to
CSV explicitly. Now it should be able
to open these CSV files.
1# Import the CSV file handler
2from typhon.files import CSV
3
4# Use the CSV file handler for the c_fileset (you could do this also
5# during initialization of the FileSet object):
6c_fileset.handler = CSV()
7
8# Maybe, the file handler needs some additional information when
9# reading a file? We can set them by *FileSet.read_args*. For example,
10# this let the file handler interpret the column 'time' as timestamp
11# object. Have a look at the CSV file handler documentation
12# to know which else parameters you can pass via read_args:
13c_fileset.read_args={
14 "parse_dates": ["time", ]
15}
16
17# This works now:
18c_fileset["2018-01-01"]
1<xarray.Dataset>
2Dimensions: (index: 15)
3Coordinates:
4 * index (index) int64 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14
5Data variables:
6 time (index) datetime64[ns] 2018-01-01 2018-01-01T00:20:00 ...
7 lat (index) float64 50.59 53.21 55.42 57.21 58.57 59.48 59.94 ...
8 lon (index) float64 -180.0 -177.5 -175.0 -172.5 -170.0 -167.5 ...
9 data (index) float64 99.7 105.6 114.1 114.6 117.0 119.8 120.1 ...
There are more file handlers for other file formats. For example,
CloudSat can read CloudSat HDF4
files. Have a look at Handlers for a complete list of official
handler classes in typhon. Every file handler might have its own specifications
and options, you can read about them in their documentations.
Handling your file formats
If you need a special format that is not covered by the official file handlers,
you can use the generic
FileHandler object and set customized
reader and writer functions. Another way - if you like object-oriented
programming - is to subclass
FileHandler and write your own file
handler class (see Create your own file handler for a tutorial). Since the latter is for
more advanced programmers, here is a simple but extensive example that shows
how to use your own reader and writer functions easily. This also shows how to
create a new fileset with many files on-the-fly:
1from datetime import datetime, timedelta
2
3# Get the base class to use a customized file handler
4from typhon.files import FileHandler
5
6
7# Here are our reader and writer functions:
8def our_reader(file_info, lineno=0):
9 """Read the nth line of a text file
10
11 Args:
12 file_info: A FileInfo object.
13 lineno: Number of the line that should be read.
14 Default is the 0th line (header).
15
16 Returns:
17 A string with the nth line
18 """
19
20 with open(file_info, "r") as file:
21 return file.readlines()[lineno]
22
23
24def our_writer(data, file_info, mode="w"):
25 """Append a text to a file
26
27 Args:
28 data: A string with content.
29 file_info: A FileInfo object.
30 mode: The writing mode. 'w' means overwriting
31 (default) and 'a' means appending.
32
33 Returns:
34 None
35 """
36
37 with open(file_info, mode) as file:
38 file.write(data)
39
40# Let's create a file handler with our functions
41our_handler = FileHandler(
42 reader=our_reader,
43 writer=our_writer,
44)
45
46# Let's create a new dataset and pass our own file handler
47our_dataset = FileSet(
48 path="data/own_dataset/{year}/{doy}/{hour}{minute}{second}.txt",
49 handler=our_handler,
50)
51
52# Fill the dataset with files covering the next two days:
53start = datetime(2018, 1, 1)
54for hour in range(0, 48, 4):
55 timestamp = start + timedelta(hours=hour)
56
57 # The content for each file:
58 text = f"Header: {timestamp}\n" \
59 + "1) First line...\n" \
60 + "2) Second line...\n" \
61 + "3) Third line...\n"
62
63 # Write the text to a file (uses the
64 # underlying our_writer function)
65 our_dataset[timestamp] = text
66
67# Read files at once and get their header line
68# (uses the underlying our_reader function)
69print(our_dataset["2018-01-01":"2018-01-03"])
1['Header: 2018-01-01 00:00:00\n', 'Header: 2018-01-01 04:00:00\n',
2 'Header: 2018-01-01 08:00:00\n', 'Header: 2018-01-01 12:00:00\n',
3 'Header: 2018-01-01 16:00:00\n', 'Header: 2018-01-01 20:00:00\n',
4 'Header: 2018-01-02 00:00:00\n', 'Header: 2018-01-02 04:00:00\n',
5 'Header: 2018-01-02 08:00:00\n', 'Header: 2018-01-02 12:00:00\n',
6 'Header: 2018-01-02 16:00:00\n', 'Header: 2018-01-02 20:00:00\n']
This script creates files containing one header line with a timestamp and some further ‘data’ lines. With our new file handler we can read easily the header line from each of those files. Great!
Pass arguments to reader and writer
The our_reader function actually provides to return the nth line of the file
if the argument lineno is given. If we want to read files with additional
arguments for the underlying reader function, we cannot use the simple
expression with brackets any longer. We have to use the more extended version
in form of the read() method instead:
1# Find the closest file to this timestamp:
2file = our_dataset.find_closest("2018-01-01")
3
4# Pass the file and the additional argument 'lineno' to the
5# underlying our_reader function:
6data = our_dataset.read(file, lineno=2)
7
8print(file, "\nData:", data)
.../data/own_dataset/2018/001/000000.txt
Start: 2018-01-01 00:00:00
End: 2018-01-01 00:00:00
Data: 2) Second line...
Using additional arguments for creating a file works very similar as above, we
can use write() here. Another
difference is that we have to generate a filename first by using
get_filename().
1# Generate a filename for our dataset and a given timestamp:
2filename = our_dataset.get_filename("2018-01-01 04:00:00")
3
4data = "4) Appended fourth line...\n"
5
6print(f"Append {data} to {filename}")
7
8# Pass the data, filename and the additional argument 'mode' to
9# the underlying our_writer function:
10our_dataset.write(data, filename, mode="a")
Append 4) Appended fourth line...
to .../data/own_dataset/2018/001/000000.txt
How can we read the second lines from all files? We could do this:
for file in our_dataset:
data = our_dataset.read(file, lineno=2)
...
If you want to use parallel workers to load the files faster (will not
make much difference for our small files here though), use
icollect() in combination with a
for-loop or simply collect() alone:
1# Read the second line of each file:
2for data in our_dataset.icollect(read_args={"lineno": 2}):
3 ...
4
5# OR
6
7# Read the second line of all files at once:
8data_list = our_dataset.collect(read_args={"lineno": 2})
Handling remote files
The FileSet class works not only for files on your local file system, but also for remote filesystems such as Amazon S3, or files in an archive such as a zip file. Please note that this functionality is still experimental and most FileHandlers only support local reading and writing, so functionality is currently limited to searching for files.
1import s3fs
2from typhon.files.fileset import FileSet
3abi_fileset = FileSet(
4 path="noaa-goes16/ABI-L1b-RadF/{year}/{doy}/{hour}/OR_ABI-L1b-RadF-M6C*_G16_s{year}{doy}{hour}{minute}{second}*_e{end_year}{end_doy}{end_hour}{end_minute}{end_second}*_c*.nc", name="abi",
5 fs=s3fs.S3FileSystem(anon=True))
6for f in abi_fileset.find("2019-11-18T05:30", "2019-11-18T07:00"):
7 print(f)
This will return all full-disk ABI L1B granules between the indicated times, for all channels. The resulting files can then be downloaded or read using the s3 interface or directly with typhon if a FileHandler is file system aware (not implemented yet). Note that unlike searching on a local file system, one should not include a leading / with the search path when searching on an s3 file system.
Get information from a file
The Dataset object needs information about each file in order to find them
properly via find(). Normally, this
happens by using placeholders in the files’
path and name. Each placeholder is represented by a regular expression that is
used to parse the filename. But sometimes this is not enough. For example, if
the filename provides not the end of the file’s time coverage but the file does
not represent a single discrete point. Let’s have a look at our Satellite C
for example:
1from typhon.files import CSV
2
3# Create a CSV file handler that interprets the column 'time' as
4# timestamp object.
5csv_handler = CSV(
6 read_csv={"parse_dates":["time", ]}
7)
8
9c_dataset = Dataset(
10 path="data/SatelliteC/{year}/{doy}/{hour}{minute}{second}.dat.gz",
11 handler=csv_handler,
12)
13
14for file in c_dataset.find("2018-01-01", "2018-01-01 8:00:00"):
15 print(file)
1.../data/SatelliteC/2018/001/000000.dat.gz
2 Start: 2018-01-01 00:00:00
3 End: 2018-01-01 00:00:00
4.../data/SatelliteC/2018/001/060000.dat.gz
5 Start: 2018-01-01 06:00:00
6 End: 2018-01-01 06:00:00
As we can see, are files interpreted as discrete files: their start time is identical with their end time. But we know that is not true, e.g. …/data/SatelliteC/2018/001/000000.dat.gz covers a period from almost six hours:
data = c_dataset.read("data/SatelliteC/2018/001/000000.dat.gz")
print("Start:", data["time"].min())
print("End:", data["time"].max())
Start: 2018-01-01 00:00:00
End: 2018-01-01 05:59:59
We have two options now:
Use the parameter time_coverage of the Dataset to specify the duration per file. Works only if each file has the same time coverage. This is the easiest and fastest option.
Using the file handler to get more information. The file handler can more than only reading or creating files in a specific format. If its method
get_info()is set, it can complement information that could not be completely retrieved from the filename.
Let’s try at first option 1:
c_dataset.time_coverage = "05:59:59 hours"
for file in c_dataset.find("2018-01-01", "2018-01-01 8:00:00"):
print(file)
1.../data/SatelliteC/2018/001/000000.dat.gz
2 Start: 2018-01-01 00:00:00
3 End: 2018-01-01 05:59:59
4.../data/SatelliteC/2018/001/060000.dat.gz
5 Start: 2018-01-01 06:00:00
6 End: 2018-01-01 11:59:59
It works! But what if each file has an individual duration? Then we need to define a file handler that have a get_info method:
TODO: The tutorial will be continued.
Placeholders
Temporal placeholders
Allowed temporal placeholders in the path argument are:
Placeholder |
Description |
Example |
|---|---|---|
year |
Four digits indicating the year. |
1999 |
year2 |
Two digits indicating the year. [1] |
58 (=2058) |
month |
Two digits indicating the month. |
09 |
day |
Two digits indicating the day. |
08 |
doy |
Three digits indicating the day of the year. |
002 |
hour |
Two digits indicating the hour. |
22 |
minute |
Two digits indicating the minute. |
58 |
second |
Two digits indicating the second. |
58 |
millisecond |
Three digits indicating the millisecond. |
999 |
All those place holders are also allowed to have the prefix end (e.g. end_year). The FileSet will use them to retrieve the start and end of the time coverage from the file path.
User-defined placeholders
Further recipes
TODO: Split this section and move it to FAQs.
Use parallel processing
If you have many files in your fileset and you want to apply a function on all
of them, you can use map() to apply this
function in parallel processes or threads.
This simple example collects the start times from all files with parallel processes:
1b_fileset = FileSet(
2 path="data/SatelliteB/{year}-{month}-{day}/"
3 "{hour}{minute}{second}-{end_hour}{end_minute}{end_second}.nc"
4)
5
6def get_start_time(file_info):
7 """Simple function to get the start time from each file"""
8 return file_info.times[0]
9
10# Collect all start times in parallel processes:
11start_times = b_fileset.map(get_start_time)
You can control which type of parallel workers is used by setting worker_type to process or thread. The numbers of workers can be set via max_workers.
How about processing the content from all files in a subroutine? For example, if we want to calculate the mean from all files. We can use the on_content option for doing this.
1def get_mean(data):
2 """Simple function to get the average of the data"""
3 return data["data"].mean()
4
5averaged_values = b_fileset.map(
6 get_mean,
7 # The on_content option passes the read content of the file instead its
8 # info object to the function:
9 on_content=True
10)
The on_content option passes the read content of the file instead of its info object to the function. If you need the file info object as well, you can set pass_info to true.
You can limit the time period that should be processed, by passing start and end:
1def get_mean(data):
2 """Simple function to get the average of the data"""
3 return data["data"].mean()
4
5averaged_values = b_fileset.map(
6 get_mean, on_content=True, start="2018-01-01", end="2018-01-02",
7)
map() always processes all files in the
given time period, waits for all results and returns them after the last worker
has finished. This might be very time-consuming for large filesets.
imap() allows to process just a chunk of
the data and to get immediate results:
for mean in b_fileset.imap(get_mean, on_content=True):
# After the first worker has finished, this will be run immediately:
print(mean)
You should consider this option if your RAM is limited.
Copy or convert files
Use move() to copy or convert files from a
fileset.
Use filters with magic indexing
TODO
Exclude or limit to time periods
TODO